Discrete Global Grid Systems with xdggs.DGGSIndex#
See also
Learn more at the xdggs documentation page.
Highlights#
xdggs provides a unified API for interacting with various Discrete Global Grid Systems (DGGS), like HEALPix or H3. It does so by parsing out grid metadata from the spatial coordinate containing the cell ids and persisting it on a “meta-index”, an index that wraps another index (most commonly a pandas index).
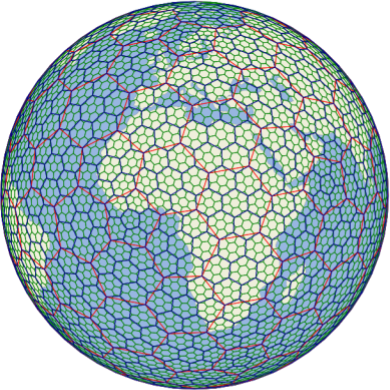
The H3 grid, a hexagonal DGGS designed for navigation on land.#
Example#
Here’s a dataset on a HEALPix grid:
import xdggs
ds = xdggs.tutorial.open_dataset("air_temperature", "healpix")
ds
<xarray.Dataset> Size: 10MB
Dimensions: (time: 2920, cells: 434)
Coordinates:
* time (time) datetime64[ns] 23kB 2013-01-01 ... 2014-12-31T18:00:00
lon (cells) float64 3kB ...
lat (cells) float64 3kB ...
cell_ids (cells) int64 3kB ...
Dimensions without coordinates: cells
Data variables:
air (time, cells) float64 10MB ...
Attributes:
Conventions: COARDS
title: 4x daily NMC reanalysis (1948)
description: Data is from NMC initialized reanalysis\n(4x/day). These a...
platform: Model
references: http://www.esrl.noaa.gov/psd/data/gridded/data.ncep.reanaly...with that, we can decode the metadata:
decoded = ds.dggs.decode()
decoded
<xarray.Dataset> Size: 10MB
Dimensions: (time: 2920, cells: 434)
Coordinates:
* time (time) datetime64[ns] 23kB 2013-01-01 ... 2014-12-31T18:00:00
lon (cells) float64 3kB ...
lat (cells) float64 3kB ...
* cell_ids (cells) int64 3kB 525 526 527 529 530 ... 2043 2044 2045 2046 2047
Dimensions without coordinates: cells
Data variables:
air (time, cells) float64 10MB ...
Indexes:
cell_ids HealpixIndex(level=4, indexing_scheme=nested, kind=pandas)
Attributes:
Conventions: COARDS
title: 4x daily NMC reanalysis (1948)
description: Data is from NMC initialized reanalysis\n(4x/day). These a...
platform: Model
references: http://www.esrl.noaa.gov/psd/data/gridded/data.ncep.reanaly...Note how the cell_ids coordinate is now associated with a HealpixIndex? This makes it harder to lose the parsed information, and thus having to parse it again.
We can also have a look at the parsed metadata:
decoded.dggs.grid_info
HealpixInfo(level=np.int64(4), ellipsoid=None, indexing_scheme='nested')
Here’s how the dataset looks:
decoded["air"].isel(time=0).dggs.explore(alpha=0.8)
In a future version of xdggs this may also support more operations, like alignment, selection using parent cell ids, or lazy coordinates for cell centers / cell boundaries.
